MIT engineers design proteins by their movement, not just their shape MIT News
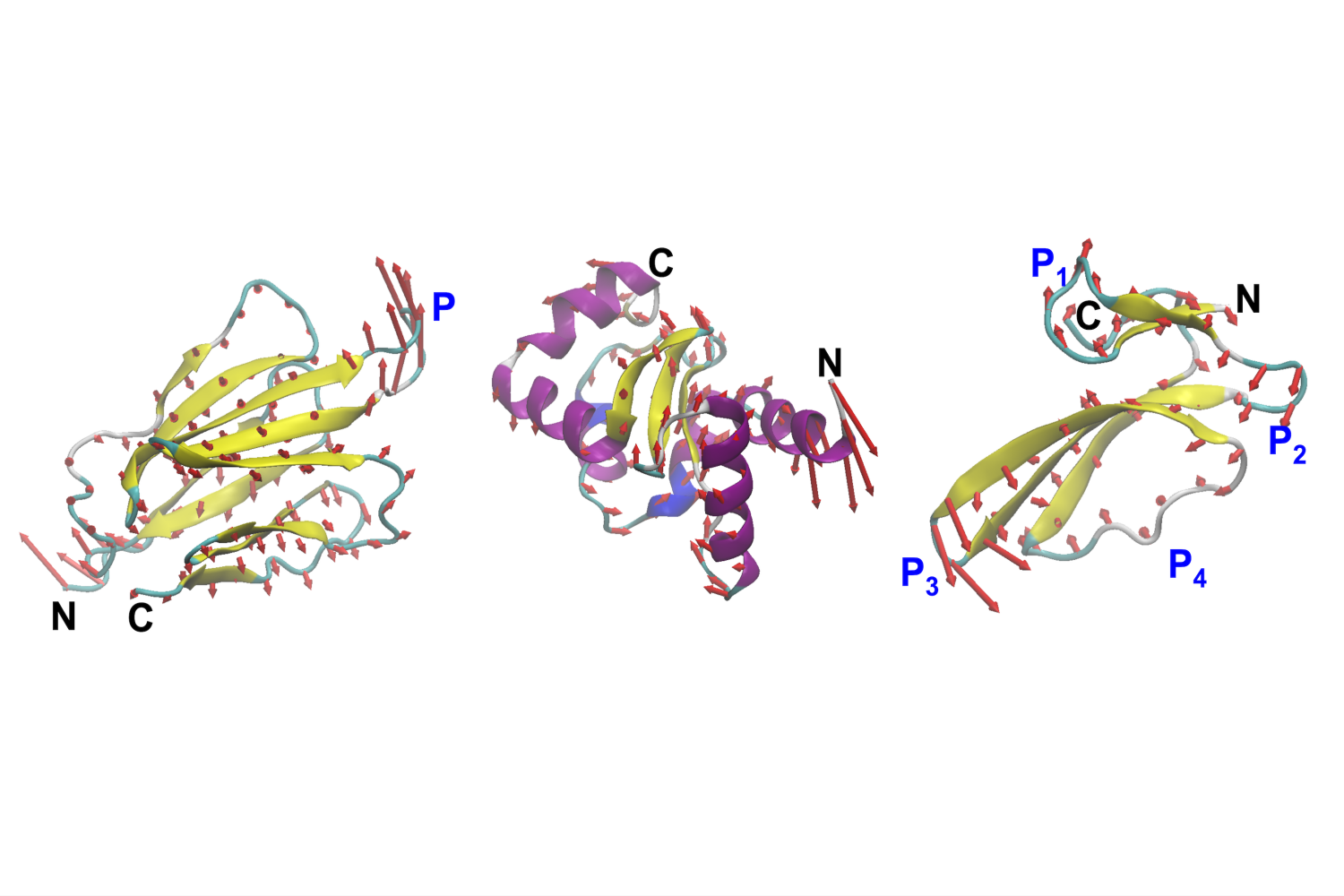
Protein is much more than the nutrients we track on a food label. Present in every cell in our body, they act as natural molecular machines. They move, stretch, bend, and contort to do their job, pumping blood, fighting disease, building muscle, and many other tasks too small for the eye to see. Their strength comes not only from the shape, but from the way they move.
In recent years, artificial intelligence has allowed scientists to design completely new protein structures not found in nature for specific functions, such as binding bacteria, or mimicking the properties of silk that work continuously. But designing a structure alone is like building a car body without controlling how the engine works. The subtle vibrations, dynamics, and mechanical dynamics of a protein are as important to its functions as its structure.
Now, MIT engineers have taken a big step to close the gap with the development of an AI model known as VibeGen. If vibe code allows programmers to define what they want and AI generates software, VibeGen does the same for living molecules: specify the vibe – the movement pattern you want – and the model writes the protein.
The new model allows scientists to understand how a protein flexes, vibrates, and moves between shapes in response to its environment, opening a new frontier in molecular engineering. VibeGen builds on a series of advances from the Buehler lab in scientific AI – systems where multiple AI models work together independently to solve the most complex problems for any one model.
“The essence of life at the basic molecular level is not just structure, but movement,” said Markus Buehler, the Jerry McAfee professor of engineering in the departments of Civil and Environmental Engineering and Mechanical Engineering. “Everything from the folding of proteins to the deformation of materials under pressure follows the basic laws of physics.”
Buehler and his former postdoc, Bo Ni, identified a critical need for what they call physics-aware AI: systems that can think in motion, not just snapshots of molecular structure. “AI must go beyond analyzing static forms to understand how structure and movement are fundamentally connected,” Buehler added.
The new method, described in a paper March 24 in the journal Important, uses generative AI to create proteins with customized dynamics.
Training AI to think in motion
The revolution in AI-driven protein science has been, surprisingly, a revolution in composition. Tools like AlphaFold have solved the decades-old problem of predicting the three-dimensional shape of a protein. Existing production models have learned to design new situations from scratch. But in focusing on the folded model — proteins frozen in place — the field ignores the area that makes proteins work: their movement. Buehler says: “Predicting the structure was so challenging that it caught the attention of the field. But the structure of a protein is one frame of a very long movie, and the design space extends in space and time, where the structure occupies a very wide area. Scientists can design a protein with a certain design. They could not yet determine how that protein would move, move, or vibrate once it was made.
VibeGen does something no protein design tool has done before. It reverses the traditional problem. Rather than asking, “What shape will this sequence produce?” it asks, “What sequence will make the protein move exactly this way?”
To build VibeGen, Buehler and Ni turned to a class of AI diffusion models, the same basic technology that powers AI image generators capable of creating realistic images out of pure sound. In the case of VibeGen, the model starts with a random sequence of amino acids and refines it, step by step, until it converges on a sequence that is predicted to vibrate and bend in a targeted way.
The system works with two interactive agents that design and challenge each other. The “designer” suggests a candidate target sequence for the target motion profile. “Prediction” examines those candidates, asking whether they will actually deliver as the designer intended. The two models iterate back and forth like an internal conversation, until the design settles into something that meets the goal. By specifying these vibrational fingerprints as design inputs, VibeGen reverses conventional thinking: dynamics becomes the blueprint, and structure follows.
“It’s a collaborative process,” Ni he says. “The designer proposes, predicts criticism, and the design evolves through that tension.”
Most of the sequences generated by VibeGen are completely de novo, not borrowed from nature, not a variation of something already done. To ensure that the designs actually work, the team performed detailed molecular physics-based simulations, and the proteins behaved exactly as intended, flexing and vibrating in the patterns that VibeGen targeted.
One of the study’s most impressive findings is that multiple protein sequences and folds can satisfy the same vibrational target — a property the researchers call functional degradation. When evolution converges on a single solution, VibeGen reveals a whole family of alternatives: proteins with different structures and sequences that move in the same way. “It suggests that nature explored a small part of what’s possible,” Buehler said. “For any given dynamic behavior, there may be a large, untapped space for effective designs.”
A new frontier in molecular engineering
Controlling protein dynamics can have different applications. In medicine, proteins can change shape when they show high energy. Many therapeutic proteins work by binding to a target molecule – a virus, a cancer cell, the wrong receptor. How well they bind often depends not only on their structure, but on how well they can adapt to the intended conditions. A motion-engineered protein can bind more precisely, reduce off-target interactions, and ultimately be a safer, more effective drug.
In materials science, which is Buehler’s area of research, the properties of machines at the molecular scale affect their performance. Biological materials such as silk and collagen derive their strength and stiffness from the coordinated movement of their molecular building blocks. Designing proteins that are rigid, flexible, or vibrate in a specific way could lead to new sustainable fibers, impact-resistant materials, or other biodegradable materials for petroleum-based plastics.
Buehler sees other possibilities: materials for buildings or cars that include protein-based components that heal themselves after mechanical stress, or repair themselves in response to heavy load.
By enabling researchers to specify motion as a direct design parameter, VibeGen treats proteins less as static shapes and more as programmable mechanical devices. This advance includes artificial intelligence, medicine, synthetic biology, and material engineering – to a future where molecular machines can be designed with the same precision and purpose as bridges, motors, or microchips.
“VibeGen can step into uncharted territory, suggesting protein designs beyond the established, perfectly designed evolutionary repertoire. It’s as if we’ve invented a new creative engine that designs molecular machinery where it’s needed,” adds Buehler.
The researchers plan to further refine the model and validate their designs in the lab. They also hope to combine motion-aware design with other AI tools, building on systems that can design proteins to be not only dynamic, but multi-functional; machines that can sense their environment, respond to signals, and adapt in real time.
The word “vibe” comes from vibration, and Buehler sees the connection as more than a play on words. “We turned the ‘vibe’ into a metaphor, a feeling, something that resonates,” he said. But in protein, vibe is physics. It is the actual pattern of motion that determines what the molecule can do, the very machinery of life.”
The research was supported by the US Department of Agriculture, the MIT-IBM Watson AI Lab, and MIT’s Generative AI Initiative.